It’s a common nightmare for programmers to come in late to a project or organization and then have to make sense of a complex “spaghetti mess” of code created over the previous 10 years — a technical debt that takes huge resources in time and money to clean up. Ten years of technical debt is an all-too common headache: Decades of debt were at the root of the Y2K COBOL nightmare. MySpace struggled famously for years with a crippling tower of technical debt. And today, both fast-growing startups and long-standing companies have to deal with legacy code on an ongoing basis in their engineering organizations and beyond.
But none of this compares to the billions of years of “technical debt” in biology. Over the last few billion years, evolution has been creating its own version of an MVP — a minimal viable product that’s shipped year after year, and works in practice, but that also never gets refactored. Somehow, it all works, because biology — at all scales, from molecular to nano to macro — takes advantage of some antifragility, the property where “the resilient resists shocks and stays the same [but] the antifragile gets better”; in spite of its complexity, it just works.
None of this compares to the billions of years of ‘technical debt’ in biology. Over the last few billion years, evolution has been creating its own version of an MVP — a minimal viable product that’s shipped year after year, and works in practice, but that also never gets refactored.
Yet in biology it also leaves a mess in its wake… and understanding that mess is the challenge that lies of ahead of us as we look to engineer improvements in our bodies, our lifespans, our health. More practically, given the rapid advances in machine learning and the new era we’re in with biology, what does healthcare innovation look like as we move beyond science to engineering with biology?
When human limitations mean we oversimplify things
The discipline of biology has, at heart, always been centered around understanding technical debt. While many biologists relish the field’s immense complexity, unfortunately, understanding that complexity is fundamentally limited by the limits of the human brain itself. We can’t even visualize let alone understand, for example, a complex mesh of thousands of interacting proteins with thousands of relevant point mutations. (That sort of complexity is unheard of in systems built by humans. I once heard a colleague of mine refer to an engineering system built with 7 degrees of freedom as the “mother of all complex human systems” — but biology tops that by many, many orders of magnitude, in even the simplest of organisms.)
We’ve oversimplified things to address our human limitations in understanding and studying complexity.
We’ve oversimplified things to address our human limitations in understanding and studying complexity. Just look at the linear, hierarchical conception of biology we teach and use: molecules in proteins, proteins in cellular machinery, machinery in cells, cells in tissue, tissue in organs, organs in organisms, organisms in niches, niches in ecosystems, ecosystems in planets, planets in solar systems, solar systems in galaxies… and so on. This hierarchy appears to convey how nature deals with evolution and change, and aids human understanding when the questions of interest are well-aligned with natural biological hierarchies. But what happens when we find something too complex, too non-linear, and too interconnected to accurately break down in this manner?
When nature itself doesn’t afford us a hierarchical structure, imposing one leads to oversimplification, which in turn limits understanding the underlying natural biological complexity. And that in turn limits the opportunities of what we can build and do with biology for better living. It’s like letting technical debt accrue to such a point that we’re missing the benefits and opportunities of understanding the complexity and refactoring the code underneath. And then there’s all the stuff we oversimplify because we don’t understand it at all (or worse, don’t know what we don’t know). Since we don’t understand the sequence of cause and effects, we may attribute breast cancer to a mutation in BRCA. But are all mutations in BRCA oncogenic, that is, leading to cancerous tumors? Often, deleterious mutations are met with compensatory mutations in other proteins to balance out the deleterious effects.
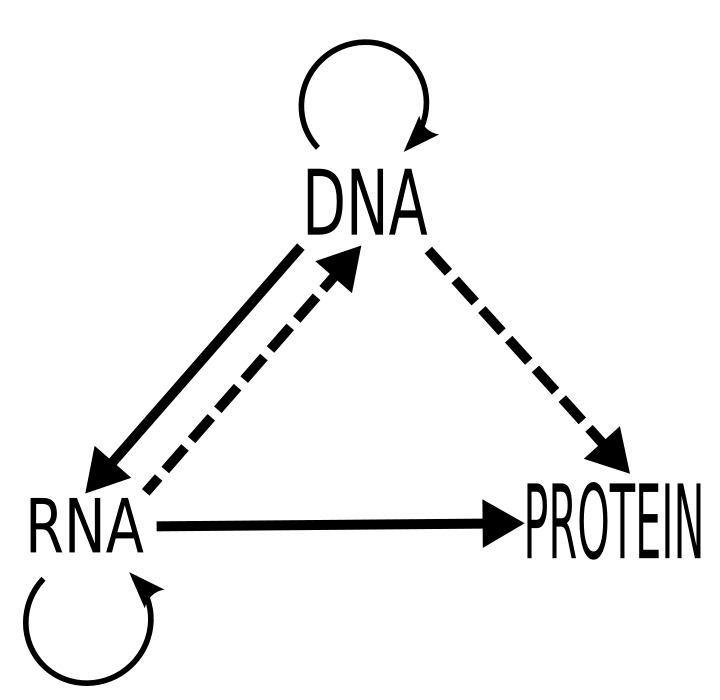
the ‘central dogma’ of biology
Even the staid old Central Dogma of Biology (DNA –> RNA –> proteins) is now understood to be far more complex, with DNA, RNA, and proteins all interacting with each other in non-linear ways. Seeing them and analyzing them in a more “high-res” way — just think about what happened when we went from analog to digital to HD! — not only gives us a clearer picture of what’s happening (especially when it comes to disease), but also enables machine learning to translate beautifully to biology. In genomics, for example, a genome is like a 1-D grid of pixels. So what happens when we get more dimensions layered into that view, thanks to advances in technology?
Machine learning — and artificial intelligence — comes in to do (and reveal) new things
The discipline of machine learning, by definition, handles the very complexity our limited human minds can’t handle. This may seem obvious on the surface: Of course a computer can handle complex calculations beyond the capability of a human! But I’m not talking about computers calculating faster and better — I’m talking about computers learning and understanding.
Machine learning in its simplest form could be described as a form of regression, or curve fitting, where a computer gets training data and then produces a predictive mathematical model based on that. Even when regressions become increasingly complex (with non-linear functions, multiple inputs, multiple dimensions, and so on), a human still needs to first understand what the key variables are — aka “features” in machine learning — so the computer can build a predictive model out of it. Without a human being giving these features to a computer, the computer would be useless, even with an infinite amount of data.
But what happens when a computer no longer needs that human input? As the fields of artificial intelligence, deep learning, and machine learning have steadily evolved over the past few decades, we may have finally turned the corner from fancy calculations to actual intelligence. New algorithms can identify and infer the key features directly from the data, sometimes even based on very limited data, taking our understanding further and even completely out of the loop. Such artificial intelligence can figure out the relevant degrees of freedom in building models, and incorporate many disparate features, all while avoiding the risk of overfitting (aka regularization). By maintaining a healthy degree of Occam’s Razor — that the simplest explanation often suffices — the system can avoid spurious conclusions.
This, by the way, is where machine learning really becomes artificial intelligence, and where deep learning comes in — it’s a matter of degree not just kind, taking the outputs from one neural network into another, in many deep layers. Deep learning has already reshaped many areas, beginning most strongly with image recognition, which is key to much of how medicine is performed. Advancements in the field have already impacted radiology, diagnosis, cancer detection, and so on — anywhere that visual pattern recognition is key — often rivaling and potentially exceeding human skills. The practical implications of this for scaling the healthcare system given lack of access and limited specialty doctors are staggering. And there is still so much exciting work being done and yet to be done in areas that are not as easily (or visually) mappable. How does one represent a drug, for example? Recent advances such as the DeepChem open source project [part of my work at Stanford University] have made strides, but it’s still only heralding what’s coming next.
So how does all this help unravel the field of biology, and go beyond the limits of the human mind and inherited tech debt? Let’s take the analogy of biomarkers that identify the presence of some disease or other change in the body. Because traditional biomarkers used in clinical tests were based on a human-limited approach to understanding the human body, they are pretty low in accuracy. But now, we can identify new biomarkers that we didn’t even know about — like for the process of aging — thanks to machine learning and AI.
When we can go beyond human hunches to really understanding biology, we get far greater predictive power. And knowing and building more accurate diagnoses and tests leads to better healthcare. A classic example is the PSA test for prostate cancer, which is only about 50% accurate, leading to a huge number of false positives. But by building on more complex features, deep learning approaches can go beyond existing and known biomarkers to explore the space, for instance, of the body’s more general immune response due to cancer. Science can’t know a priori, based on the limits of human research and understanding, what such an immune response would like look — but a deep learning algorithm can, learning it for the very first time. This same phenomenon plays out across machine and deep learning applied to other medical issues as well, from heart disease and beyond.
When we can go beyond human hunches to really understanding biology, we get far greater predictive power.
Welcome to a new era of engineering biology
For the first time, the technical debt and “spaghetti code” of biology can be mapped, understood, and even refactored. And given the better-than-Moore’s-Law for bio, this is happening at a time when genomics, proteomics, metabolomics, etc. have become relatively inexpensive to map. Coupled with the advances in AI (which itself are driven by similar cost reduction curves), this all opens the door to new applications of biology for healthcare with unprecedented accuracy.
So the question becomes: When you finally understand the spaghetti code of bio, what can you do with it? The answer is that we’re living in a new age of science, where empiricism is being replaced by engineering. What do I mean by that? Think about a bridge, big or small, that you cross on your way to work everyday or that you once saw on vacation: It was, as all bridges are, engineered, because we know exactly how to build them, and build them well. But in many areas in biology, we don’t yet know how to build things, so instead of engineering, we discover them more empirically (hypothesize, test, analyze, conclude). Once upon a time, bridges were built that way too — in the Roman era, some bridge experiments failed, some lasted; we studied why, and we learned. That’s empiricism.
And that’s the phase of biology we’ve been in until now. There’s a reason pharmaceutical development is still called “drug discovery”; it involves lots of empirical testing of many, many variations to see which one will work, being tested from tubes to animals to people. But once you understand more of the spaghetti code here, you not only get faster tests (high throughput screening) of more diversity in molecules (combinatorial chemistry) for drug discovery, you can also rip it all out and start anew.
Just as programmers prefer to start from scratch when it comes to technical debt, biologists and developers can now, too, build entirely new things from scratch. Imagine being able to engineer biological circuits with the sophistication of evolution, yet without the technical debt. It’s been considered impossible until now, but recent biochemical engineering advances have created a set of minimally interacting proteins that allow new circuits to take advantage of useful cellular machinery without disrupting them. The next challenge, of course, is how to engineer those circuits directionally instead of just empirically testing a bunch of possible circuits; that’s where electronic design automation (EDA) software for cells comes in. These are the building blocks for designing with biology — using software engineering principles of abstraction, modularity, and hierarchy — and point to what’s possible when we move from science to engineering. It is, quite frankly, the beginning of truly “intelligent design”.
So what are the practical and immediate implications of all this? Here are a few quick examples: Instead of classic, costly, clinical trials, pharmaceutical companies can now replace theory with data itself to advance engineering. Meanwhile, with digital therapeutics, patients and doctors can achieve not only better efficacy of treatment for diseases like diabetes, but also constantly improve that efficacy in a way that could never happen with a drug. (It’s like A/B testing, without the clinical trials, and without any toxicity whatsoever.) Perhaps most interestingly — especially given the confusing legacy that is our current healthcare system — engineering principles can be applied to the health care system as a whole. I’d argue that many of the problems we face here are just another type of “technical debt”: a tangled weave of political, medical, and financial factors given how the U.S. healthcare system has evolved. While starting from scratch here is clearly unrealistic, given the vast network of functions that must remain functional, a new set of startups can work within the system to provide much needed transparency and information while decreasing costs.
And we can go even further. Our entire health system is centered around paying for treatment of specific disease indications; what happens when we shift mindsets from what to identify to what to prevent? If you really think about it, healthcare should really be about stopping us all from getting sick and preventing harm in the first place — not just the current paradigm of “do no harm”. Admittedly, “prevention” is a loaded term, which brings to mind all the things we keep hearing we’re supposed to do, or tests that are too low accuracy to help. We can break those legacy barriers with more accurate tests (and without false positives taxing the system), as well as with tests that we can really use (and that have high impact outcomes).
Healthcare should really be about stopping us all from getting sick and preventing harm in the first place — not just the current paradigm of ‘do no harm’.
But beyond a systemic level, what does all this mean for individual doctors? For one thing, we’re now entering an era where the seemingly helpful Hippocratic oath that doctors “take” needs to be turned on its head. Because in the end, true healthcare transformation will come from shifting mindsets from “do no harm”/ “don’t kill” to “doing right” — that is, “prevent” and “heal”. Ensuring people are healthy should not only be about treating disease! (Especially since healthy people are generally outside of the medical system until they develop a disease.) We are finally poised to move away from such a hit-or-miss model — and it’s really a shift away from the lucky hunches of empiricism towards the predictability of engineering biology.
None of this, I believe, will take people — and especially doctors — completely out of the loop. The true winner in the end will be augmented intelligence. Just as humans built other tools to push ourselves forward — the x-ray, the stethoscope, the EKG, the smartphone — this too is another “bicycle for our minds”. Thinking in terms of augmentation is, I believe, the key to unlocking and understanding biology’s technical debt. It’s that attitude that will carry us forward to a truly brave new world, one where machines help make us better humans.
images: central dogma via Wikimedia Commons
