Our best tools for understanding and manipulating biology have historically been observation and experimentation. Then, we developed the ability to “read” and “write” in biology. And now, the convergence of biology with computation and engineering has finally unlocked our ability to “execute” in biology.
In other words: biology is becoming programmable.
Read (r)
Each time our tools advance, from observing nature with our naked eye to the invention of nano-microscopy, we see biology more clearly and completely. Today we don’t just see; we can also read across the many programming languages biology uses: genomics (DNA); transcriptomics (RNA); epigenomics (genome structure); proteomics (proteins); metabolomics (metabolites); microbiomics (resident bacteria). Reading with this multi-omics approach enables us to capture the full breadth of biology, and at a resolution that’s increasing every day—down to the single cell! We can see errors, interactions, and processes we have never seen before: even, for example, how each individual cell is behaving in the dynamic dance between invading cancer cells and protective immune cells.
But reading code is not the same as understanding how biology works. Biology, of course, is much more complex than any human-designed system; it’s a hairball of entangled networks that communicate, govern, and respond to each other in ways that we don’t fully comprehend. If you printed the human genome (and someone did!), those 3 billion letters come to ~130 volumes of encyclopedias. To complicate matters further, these 3 billion letters code for “only” 20,000 genes—which contain instructions for millions of variations of proteins, some of which regulate which combination of genes are turned on or off… all of which, ultimately, drives the genetic program that determines a cell’s state and fate. How do we make sense of all of that?
This is where advanced computational techniques like artificial intelligence come in. These technologies are particularly well-suited to connect dots humans wouldn’t be able to connect unaided. By combining readouts from all these different “languages,” algorithms can pick up subtle patterns correlated with diseases like cancer, or even aging. Identifying these patterns helps deepen our understanding of the governing rules for biology—which means we are now moving from sight to insight.
Write (w)
Historically we had blunt tools to write to biology, and a limited understanding of how biology works, which made it exceedingly difficult to make biology work for us. Humans have a long history of attempting to control biology—cross-pollinating crops, breeding livestock, even testing natural products that alleviate disease. But our approach was mostly experimental, and usually required brute force: we tried something, then would see what happened; if it worked, we would try to make it better. In other words, hypothesis -> test -> evaluate; revise hypothesis -> re-test -> re-evaluate.
But as we developed new ways to manipulate biology directly, we began to have the opportunity to fix certain errors we found. Four decades ago, the rise of recombinant DNA technology—the ability to splice in human genes into other organisms—birthed the biotech industry. Our next major advance was gene therapy, where a corrected copy of a gene could be delivered into a human cell to supplant a broken gene. Gene therapies have been approved for devastating diseases including congenital blindness, somatic muscular atrophy, and beta thalassemia, with many more in the pipeline. We’re now in the middle of another great advance: the ability to precisely edit the errors in existing genes directly, with CRISPR (early data is showing promise in sickle cell disease, for example). Today, with the precision of CRISPR tools and synthetic DNA advances, we can write to biology.
The question is: do we know what we want to say?
Execute (x)
The confluence of enhanced biological knowledge with advanced engineering capabilities is driving our ability to execute programs in biology. As we can increasingly analyze and understand biology, we’re gaining the biological equivalents of software debuggers and compilers. With engineering, we can now make modular components (sensors, switches, logic gates) we can stockpile, reuse, and repurpose like Lego blocks—a sort of biological code repository. And like any engineered process based on a design -> build -> test cycle, advances in programming biology will accrue, compound, and accelerate over time.
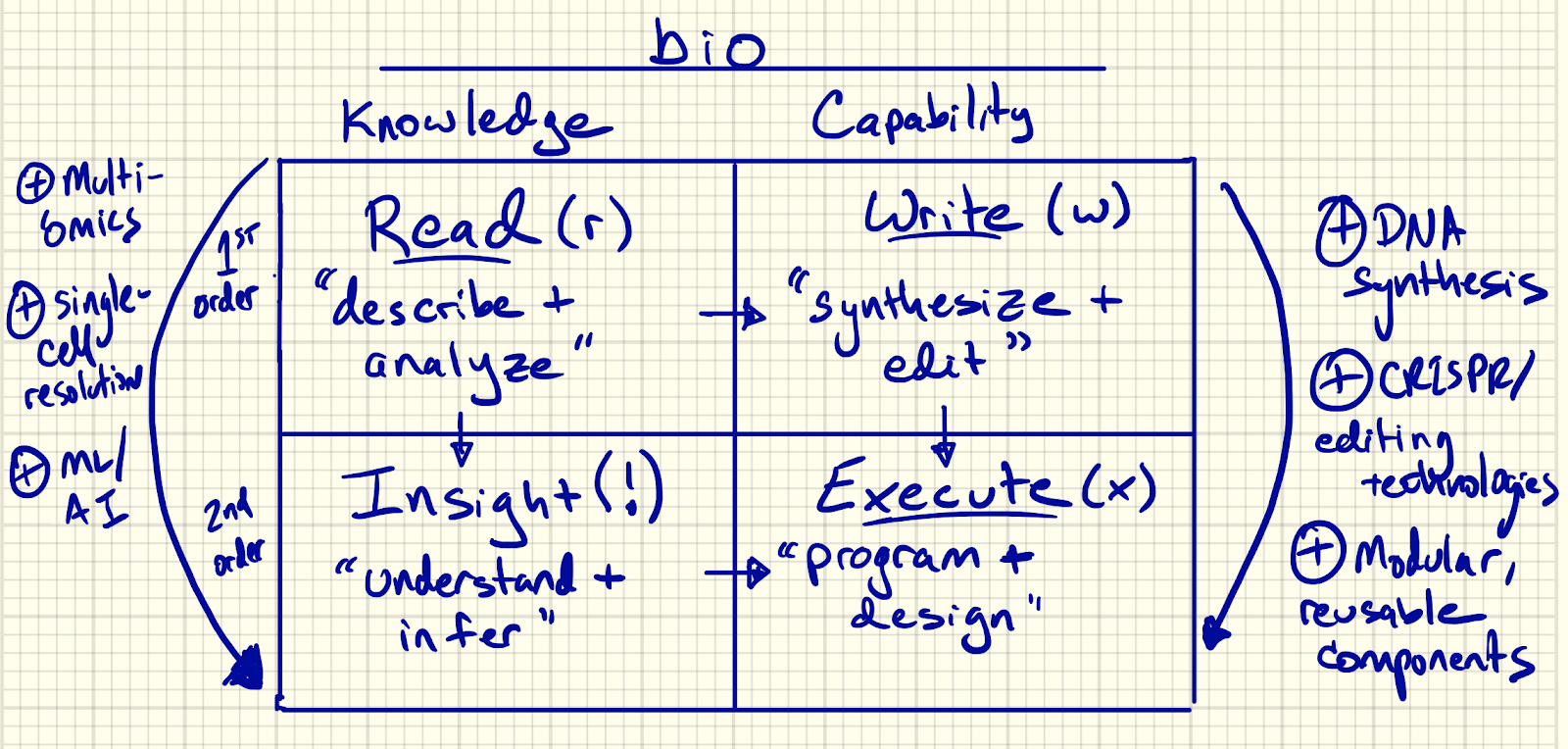
Read (r), write (w), and execute (x) in biology are inextricably linked; in many ways, they reinforce each other. It’s a knowledge-capability matrix relationship (see below) where our ability to read informs what we can write; the arrival of new technologies like AI elevates reading to insight; and the joining of writing with insight powers our ability to execute.
With its ability to store information, replicate and evolve, biology is one of the most advanced technologies on earth. Just the way software has transformed virtually every aspect of our world, this new ability to execute programs in biology will touch all of our lives, both in terms of our health and far beyond—with engineered biology applications in agriculture, food, textiles, manufacturing… even computers.
Imagine an engineered cell that can look for the presence of disease, run a computation to determine what program to run—maybe to attack an unhealthy cell, or recruit other immune cells—and then shut itself off when the job is done. CAR T is an early example of the promise for sophisticated cell therapies that do just that, to tackle complex diseases like cancer. As we become increasingly capable of executing programs in biology, we won’t be limited to just repair what’s broken, we’ll be able to use biology as a true design medium to wildly reimagine and remake the physical world around us.
Medicine is becoming programmable; and biology is eating the world.